LaTeX templates and examples — Preprints
Seneste

A template for submitting articles to paleorXiv.

A sample template to submit papers to MarXiv: the free research repository for the ocean and marine-climate sciences. Documentation for MarXiv is available at https://www.marxiv.org. The repository is located at https://osf.io/preprints/marxiv. This template is based on the engrXiv template, accessible at https://www.overleaf.com/latex/templates/engrxiv-template/ttrnvgdkgcgy.

This is a template for scientific manuscripts. It follows standard conventions for organizing a manuscript, (Abstract / Intro / Results / Discussion / Methods). Things that make it unusually nice: it uses cleverref package and "phantom" subpanels so you can refer to "Figure 1A" as a hyperlink. You can easily change to: two-column format, double/single spacing, line numbers. I work at NYU Langone Health; the "Highlight" and "Accent" colors are from the institutional style guide for NYU Langone Health's violet / blue. NYU's style guide recommended Montserrat as a web-friendly sans serif font. License is CC0 -- i.e. no restrictions. I hope you find it useful. I will not provide any support for this template.

a basic arXiv template.
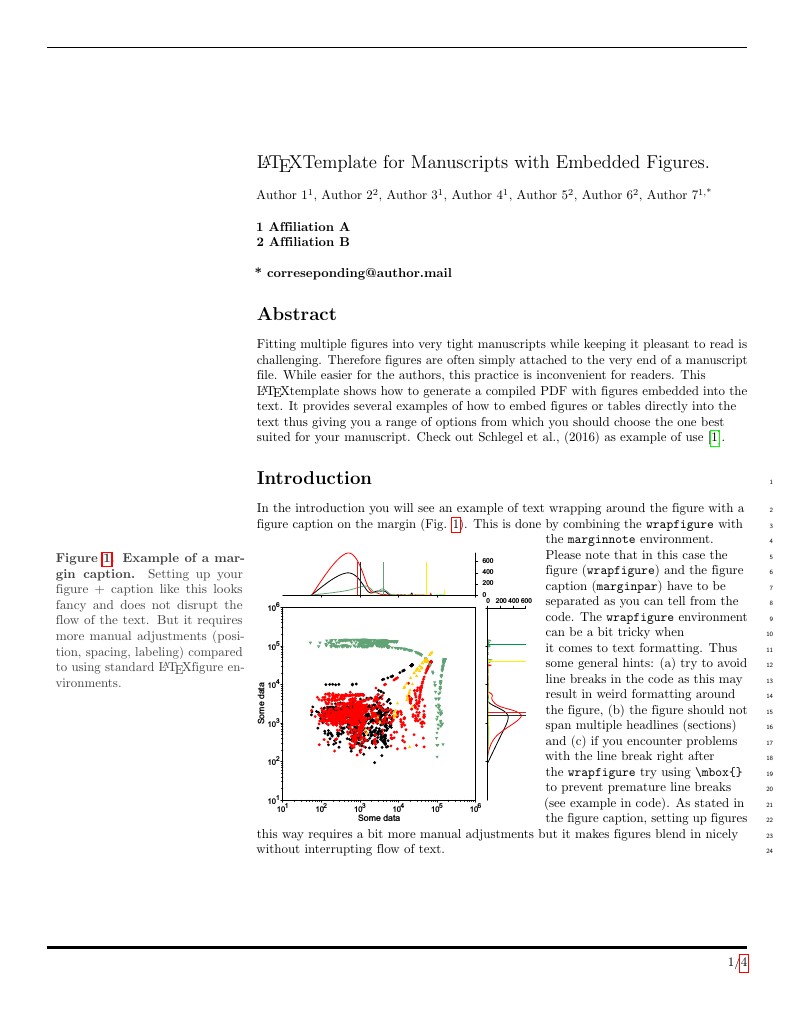
Template for writing scientific manuscripts. Features several examples of how to embed figures directly into the text. Use it to create compiled PDFs - e.g. for pre-peer review publication on the arXiv, bioRxiv, or to a repository such as figshare. To submit your manuscript to the arXiv, bioRxiv, figshare or one of many other destinations linked to from Overleaf, simply click the 'Journals & Services' button on the top bar of the Overleaf editor and choose the appropriate destination from the menu. You can also use the 'Download as zip - for submission' option in the Project menu to download a zip file containing all the required files for the submission (e.g. including the .bbl file if you've used a bibliography file for your references).

Developing a fast a versatile algorithm for curve fitting with neural networks.

LaTeX style template suitable for “preprint” publications such as arXiv and bio-arXiv.

A paper template for submissions to engrXiv, the eprint server for engineering.

This is a basic article template for MIT CSAIL preprints (i.e., submissions to arXiv).
\begin
Discover why over 25 million people worldwide trust Overleaf with their work.